New innovations to propel your research
The neXt generation of single cell starts with GEM-X
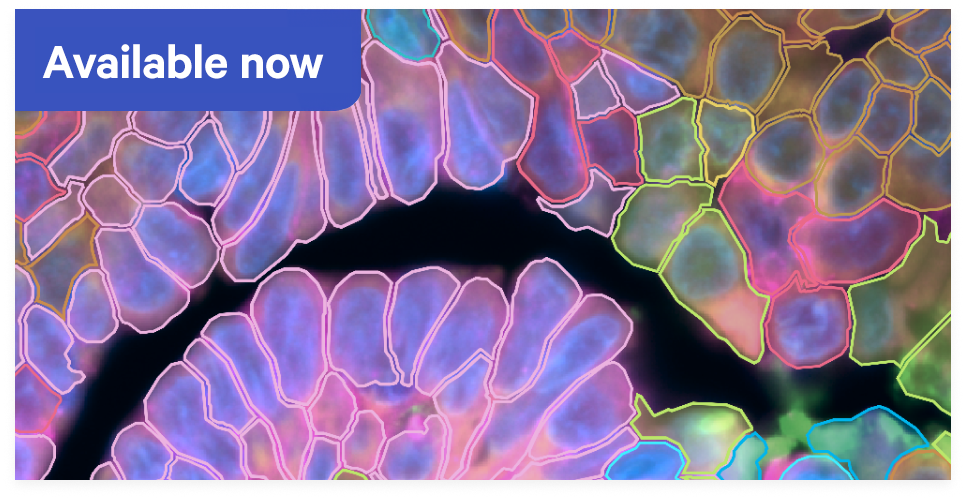
Fuel your success with enhanced performance, including increased sensitivity, robustness & throughput.A new standard for Xenium cell segmentation
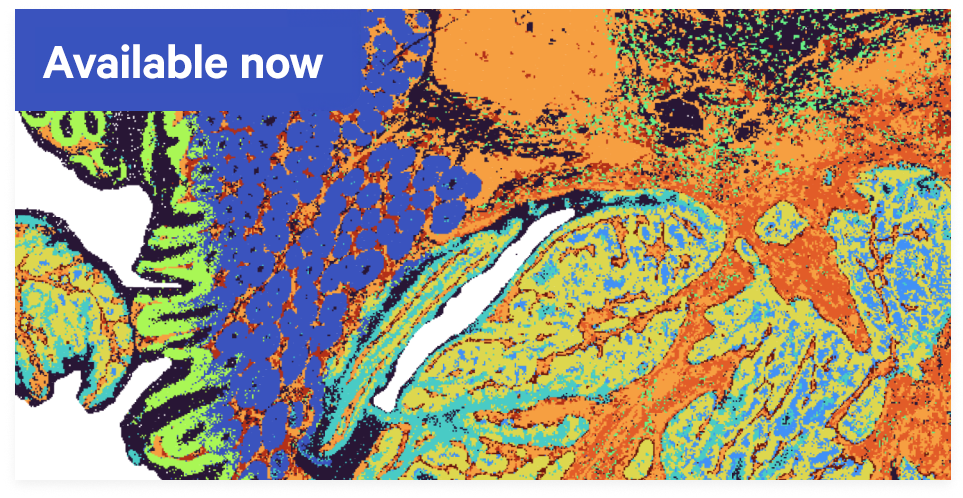
Confidently assign transcripts to cells with precise multimodal segmentation and our advanced AI algorithm.Discover the whole transcriptome in high definition
The spatial discovery power you want now with single cell-scale resolution and continuous tissue coverage.
Your toolbox to resolve biology's complexities

Chromium Single Cell
Our single cell platform allows transcriptomic profiling of up to a million single cells with multiomic capabilities to reveal cellular diversity.Learn about Chromium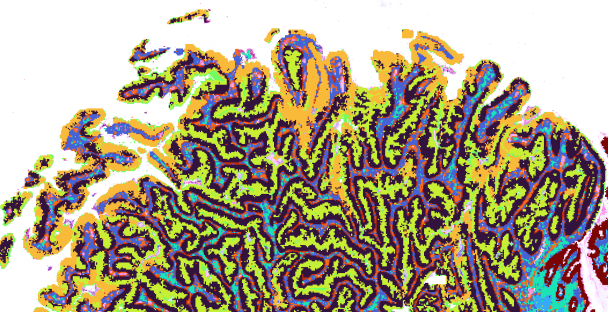
Visium Spatial
Our spatial discovery platform enables whole transcriptome analysis of FFPE & fresh frozen tissue within morphological context.Learn about Visium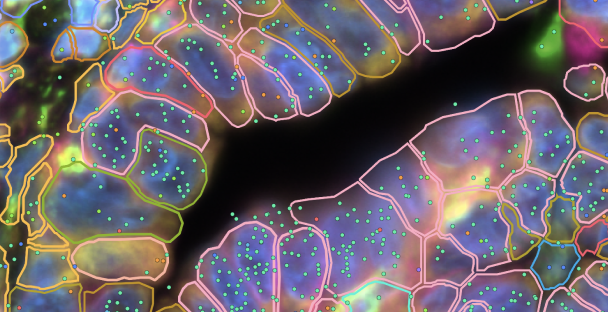
Xenium In Situ
Our spatial imaging platform maps RNA molecules with subcellular resolution in tissue sections for a deeper look into cellular mechanisms & interactions.Learn about Xenium
Our products empower impactful science
With over 7,200 publications using our products, join the community of scientists advancing their research with the aid of 10x Genomics.
Solutions to see biology in new ways
Single Cell Gene Expression
Measure gene activity on a cell-by-cell basis and characterize cell populations, cell types, and more.
Single Cell Gene Expression Flex
Perform sensitive whole transcriptome gene expression on fresh or fixed samples (including FFPE).
Single Cell Multiome ATAC + Gene Expression
Measure gene expression and open chromatin simultaneously from the same cell, across thousands of cells.
Single Cell Immune Profiling
Measure the activity of immune cells and their targets.
HD Spatial Gene Expression
Perform sequencing-based spatial transcriptomics at single cell scale with continuous tissue coverage.
In Situ Gene Expression
Spatially map gene expression at subcellular resolution with an imaging-based readout.